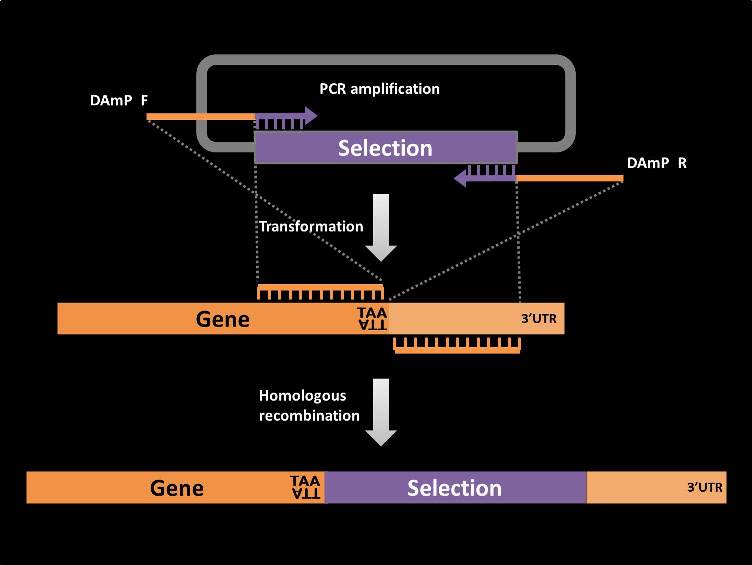
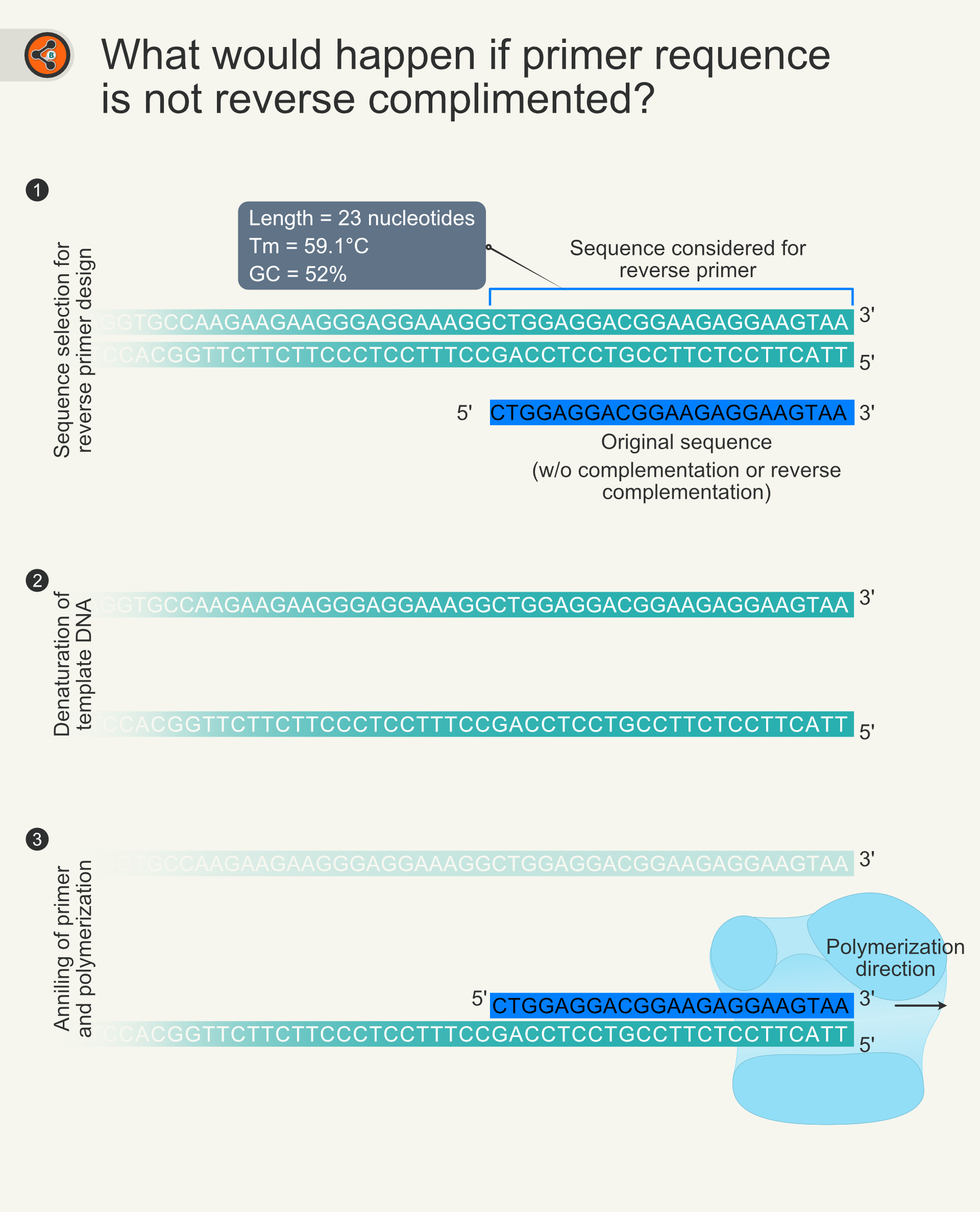
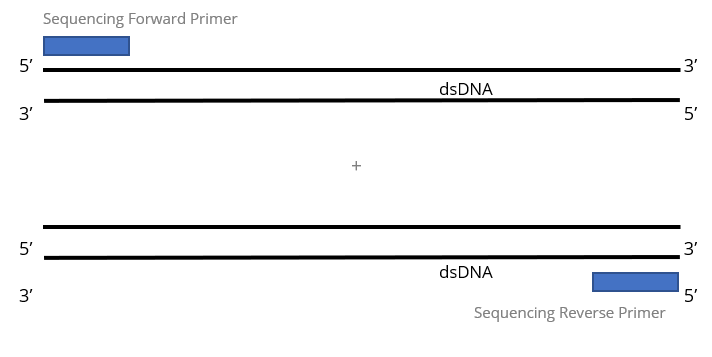
Highly specific real-time quantification of diverse microRNAs in human samples using universal primer set frame - ScienceDirect
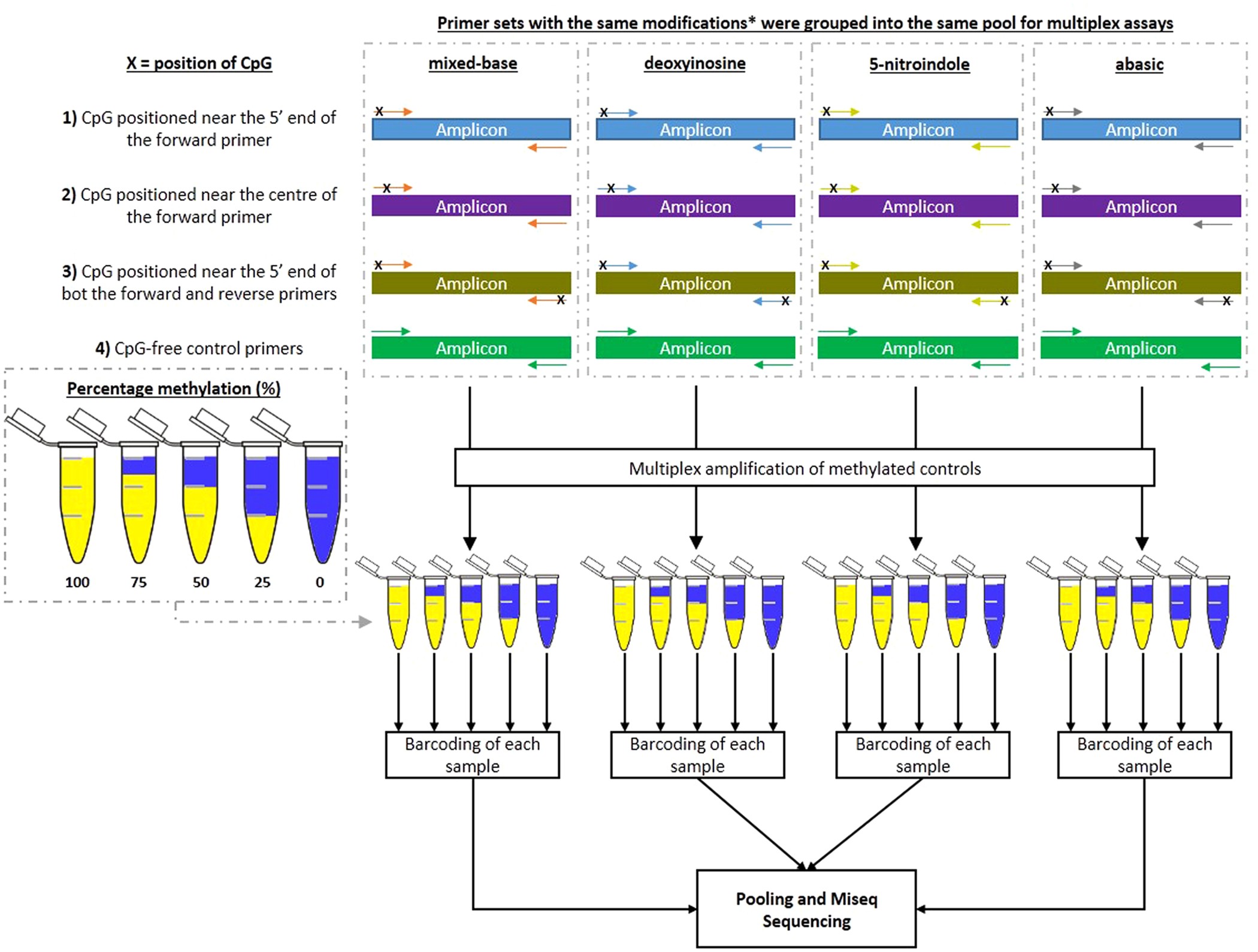
Evaluation of Different Oligonucleotide Base Substitutions at CpG Binding sites in Multiplex Bisulfite-PCR sequencing | Scientific Reports
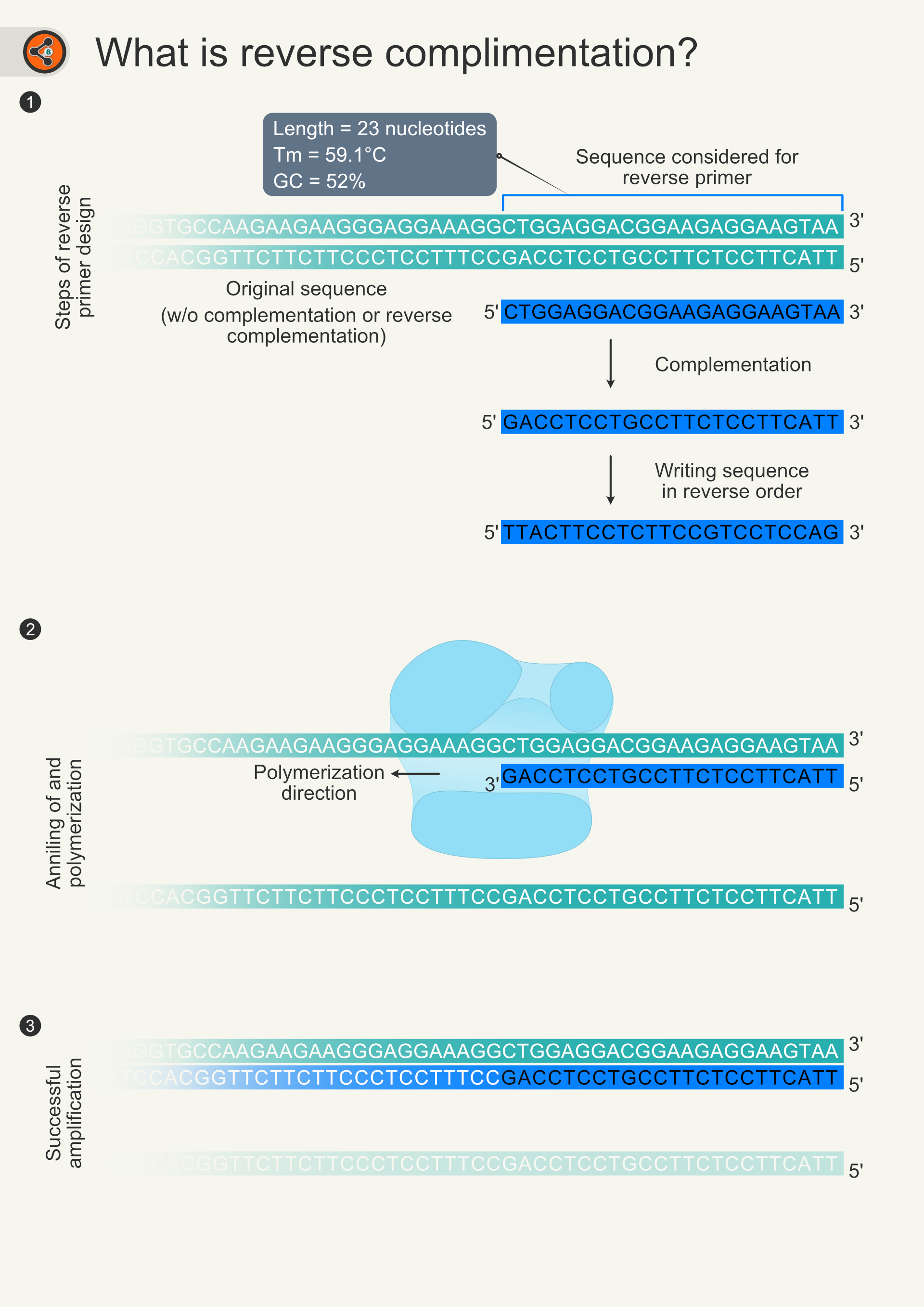
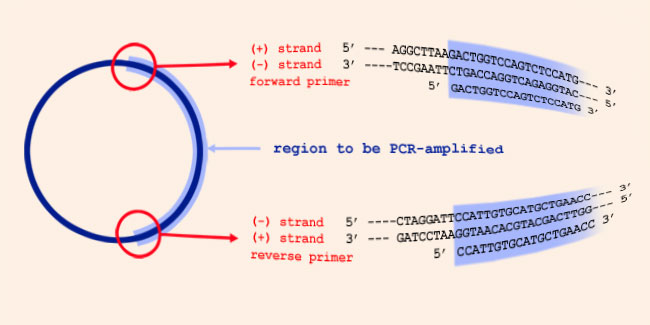
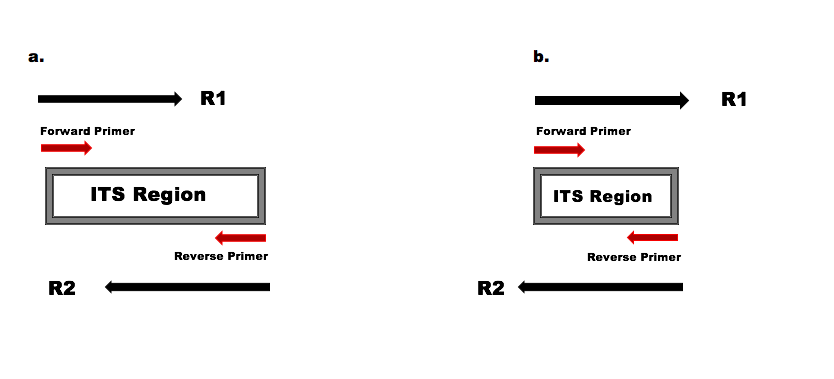
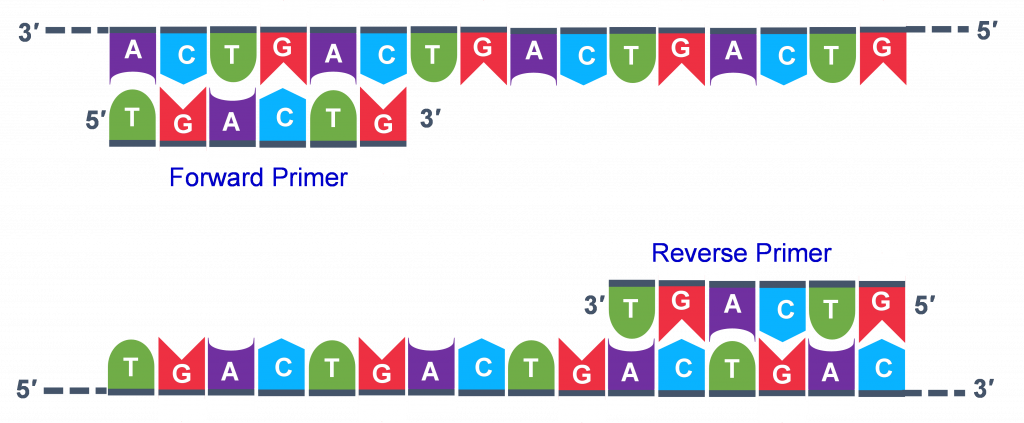
Forward and reverse primers are complementary to different DNA strands. These DNA strands are complementary to each other. Which statement is right? - Quora
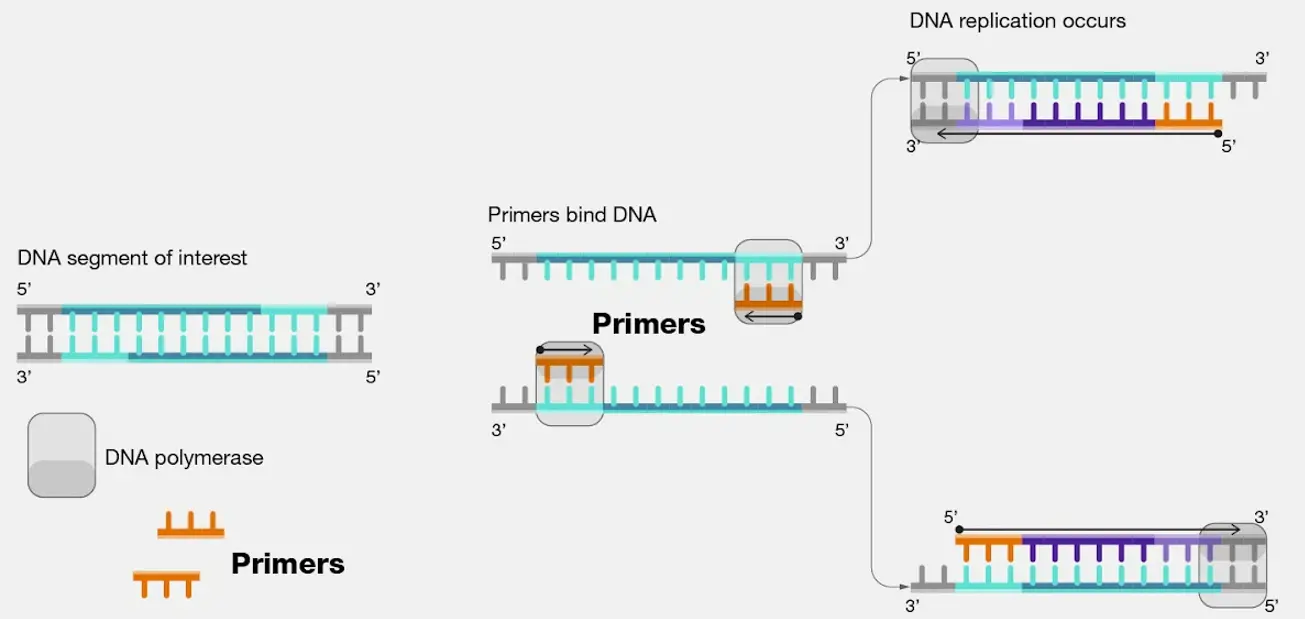
Designing PCR Primers using Primer3, UCSC in-Silico PCR and primer-BLAST Primers are short sequences of single stranded DNA that
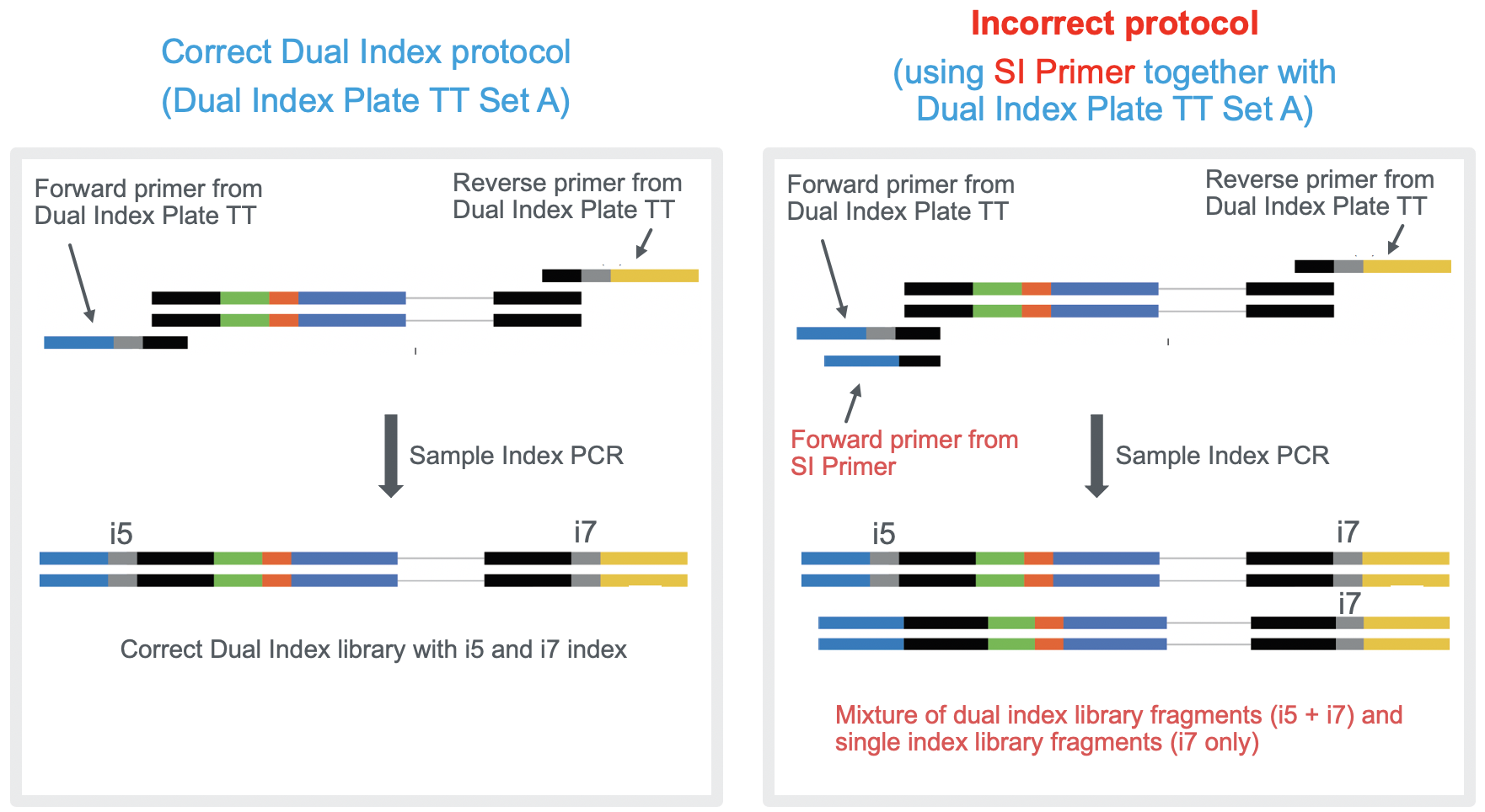
Can I use the SI Primer together with Dual Index Plate TT when preparing Single Cell 3' Gene Expression libraries? – 10X Genomics
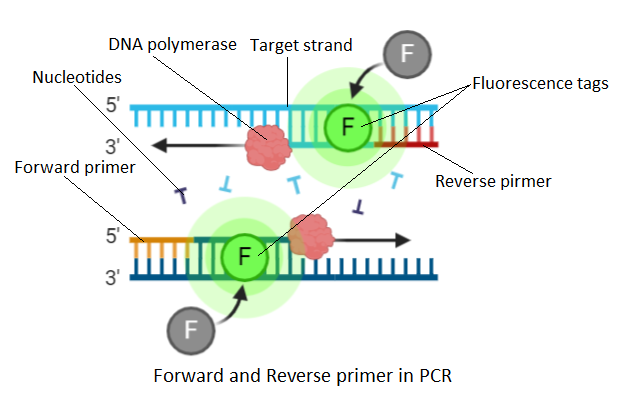
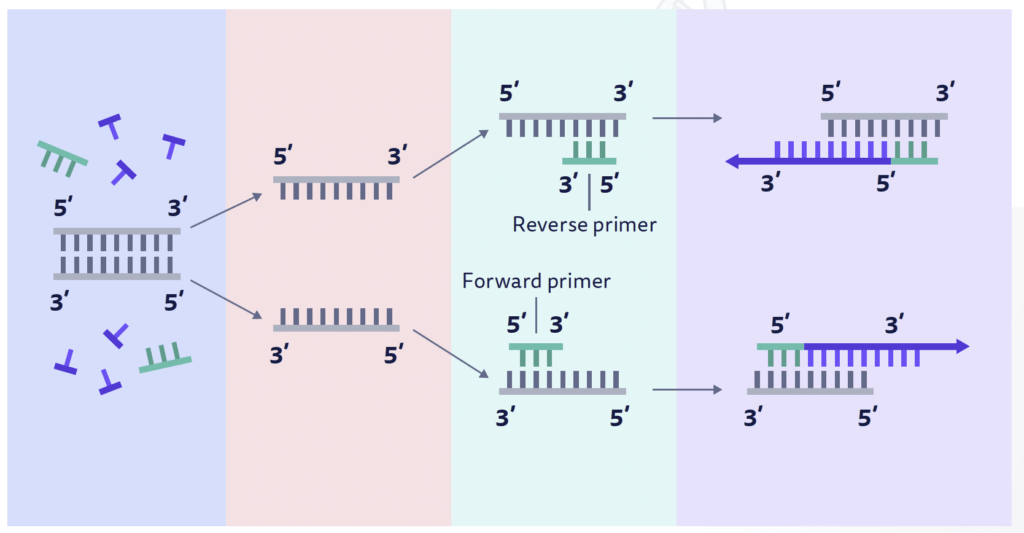
Primers (forward and reverse) are synthetic oligonucleotides of 17-30 nucleotides. They are complementary to the sequence present on the desired DNA segment. Why?
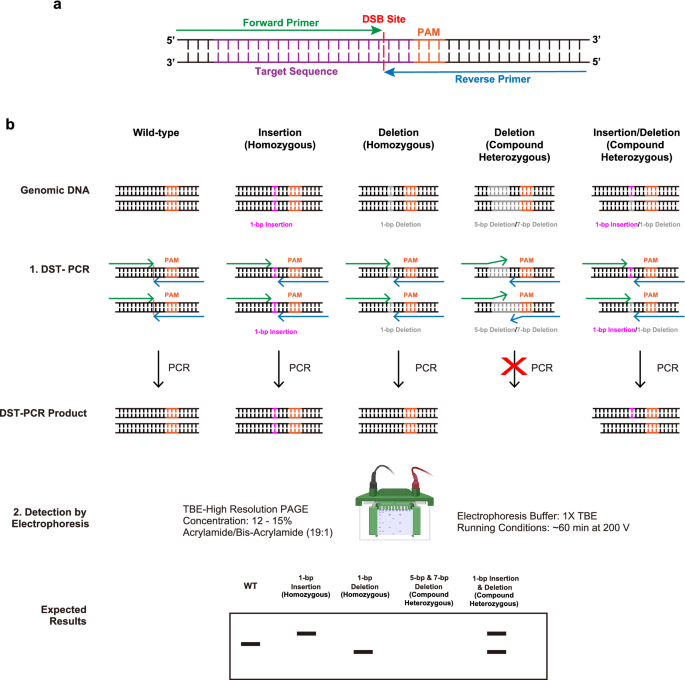
A pair of primers facing at the double-strand break site enables to detect NHEJ-mediated indel mutations at a 1-bp resolution | Scientific Reports
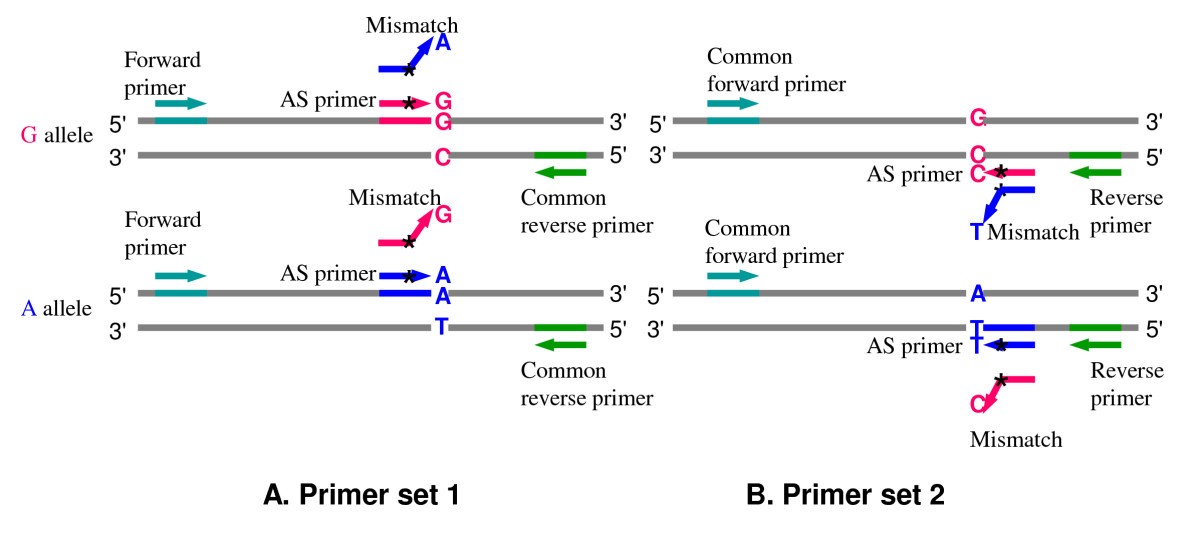
BatchPrimer3: A high throughput web application for PCR and sequencing primer design | BMC Bioinformatics | Full Text